Assessing Read Quality
Last updated on 2024-11-17 | Edit this page
Estimated time: 50 minutes
Overview
Questions
- How can I describe the quality of my data?
Objectives
- Explain how a FASTQ file encodes per-base quality scores.
- Interpret a FastQC plot summarizing per-base quality across all reads.
- Use
forloops to automate operations on multiple files.
This lesson has been adapted from the original Data Carpentry - Wrangling Genomics to be run using the NeSI infrastructure as part of the Otago Bioinformatics Spring School instead of AWS.
Bioinformatic workflows
When working with high-throughput sequencing data, the raw reads you get off of the sequencer will need to pass through a number of different tools in order to generate your final desired output. The execution of this set of tools in a specified order is commonly referred to as a workflow or a pipeline.
An example of the workflow we will be using for our variant calling analysis is provided below with a brief description of each step.
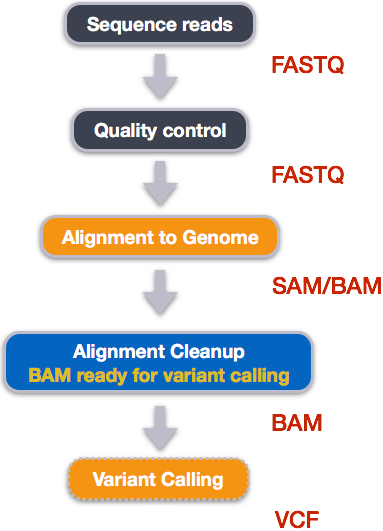
- Quality control - Assessing quality using FastQC
- Quality control - Trimming and/or filtering reads (if necessary)
- Align reads to reference genome
- Perform post-alignment clean-up
- Variant calling
These workflows in bioinformatics adopt a plug-and-play approach in that the output of one tool can be easily used as input to another tool without any extensive configuration. Having standards for data formats is what makes this feasible. Standards ensure that data is stored in a way that is generally accepted and agreed upon within the community. The tools that are used to analyze data at different stages of the workflow are therefore built under the assumption that the data will be provided in a specific format.
Starting with data
Often times, the first step in a bioinformatic workflow is getting the data you want to work with onto a computer where you can work with it. If you have outsourced sequencing of your data, the sequencing center will usually provide you with a link that you can use to download your data. Today we will be working with publicly available sequencing data.
We are studying a population of Escherichia coli (designated Ara-3), which were propagated for more than 50,000 generations in a glucose-limited minimal medium. We will be working with three samples from this experiment, one from 5,000 generations, one from 15,000 generations, and one from 50,000 generations. The population changed substantially during the course of the experiment, and we will be exploring how with our variant calling workflow.
The data are paired-end, so we will download two files for each sample. We will use the European Nucleotide Archive to get our data. The ENA “provides a comprehensive record of the world’s nucleotide sequencing information, covering raw sequencing data, sequence assembly information and functional annotation.” The ENA also provides sequencing data in the fastq format, an important format for sequencing reads that we will be learning about today.
To save time, the data has already been downloaded for you and placed
in ~/obss_2024/genomic_dna/data/untrimmed_fastq.
Today we’re going to be working inside of
~/obss_2024/genomic_dna. The main directory structure for
genomic_dna/ (inside of ~/obss_2024/)
currently looks like this currently
Now lets navigate to where the untrimmed fastq data is stored.
To download the data, run the commands below.
Here we are using the -p option for mkdir.
This option allows mkdir to create the new directory, even
if one of the parent directories does not already exist. It also
supresses errors if the directory already exists, without overwriting
that directory.
It will take about 15 minutes to download the files.
Original way to obtain the data (Use if not on NeSI)
To download the data, run the commands below.
Here we are using the -p option for mkdir.
This option allows mkdir to create the new directory, even
if one of the parent directories doesn’t already exist. It also
supresses errors if the directory already exists, without overwriting
that directory.
It will take about 15 minutes to download the files.
BASH
mkdir -p ~/obss_2024/data/untrimmed_fastq/
cd ~/obss_2024/data/untrimmed_fastq
curl -O ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR258/004/SRR2589044/SRR2589044_1.fastq.gz
curl -O ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR258/004/SRR2589044/SRR2589044_2.fastq.gz
curl -O ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR258/003/SRR2584863/SRR2584863_1.fastq.gz
curl -O ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR258/003/SRR2584863/SRR2584863_2.fastq.gz
curl -O ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR258/006/SRR2584866/SRR2584866_1.fastq.gz
curl -O ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR258/006/SRR2584866/SRR2584866_2.fastq.gzThe data comes in a compressed format, which is why there is a
.gz at the end of the file names. This makes it faster to
transfer, and allows it to take up less space on our computer. Let’s
unzip one of the files so that we can look at the fastq format.
Quality control
We will now assess the quality of the sequence reads contained in our fastq files.
Details on the FASTQ format
Although it looks complicated (and it is), we can understand the fastq format with a little decoding. Some rules about the format include…
| Line | Description |
|---|---|
| 1 | Always begins with ‘@’ and then information about the read |
| 2 | The actual DNA sequence |
| 3 | Always begins with a ‘+’ and sometimes the same info in line 1 |
| 4 | Has a string of characters which represent the quality scores; must have same number of characters as line 2 |
We can view the first complete read in one of the files our dataset
by using head to look at the first four lines.
OUTPUT
@SRR2584863.1 HWI-ST957:244:H73TDADXX:1:1101:4712:2181/1
TTCACATCCTGACCATTCAGTTGAGCAAAATAGTTCTTCAGTGCCTGTTTAACCGAGTCACGCAGGGGTTTTTGGGTTACCTGATCCTGAGAGTTAACGGTAGAAACGGTCAGTACGTCAGAATTTACGCGTTGTTCGAACATAGTTCTG
+
CCCFFFFFGHHHHJIJJJJIJJJIIJJJJIIIJJGFIIIJEDDFEGGJIFHHJIJJDECCGGEGIIJFHFFFACD:BBBDDACCCCAA@@CA@C>C3>@5(8&>C:9?8+89<4(:83825C(:A#########################Line 4 shows the quality for each nucleotide in the read. Quality is interpreted as the probability of an incorrect base call (e.g. 1 in 10) or, equivalently, the base call accuracy (e.g. 90%). To make it possible to line up each individual nucleotide with its quality score, the numerical score is converted into a code where each individual character represents the numerical quality score for an individual nucleotide. For example, in the line above, the quality score line is:
OUTPUT
CCCFFFFFGHHHHJIJJJJIJJJIIJJJJIIIJJGFIIIJEDDFEGGJIFHHJIJJDECCGGEGIIJFHFFFACD:BBBDDACCCCAA@@CA@C>C3>@5(8&>C:9?8+89<4(:83825C(:A#########################The numerical value assigned to each of these characters depends on the sequencing platform that generated the reads. The sequencing machine used to generate our data uses the standard Sanger quality PHRED score encoding, using Illumina version 1.8 onwards. Each character is assigned a quality score between 0 and 41 as shown in the chart below.
OUTPUT
Quality encoding: !"#$%&'()*+,-./0123456789:;<=>?@ABCDEFGHIJ
| | | | |
Quality score: 01........11........21........31........41Each quality score represents the probability that the corresponding nucleotide call is incorrect. This quality score is logarithmically based, so a quality score of 10 reflects a base call accuracy of 90%, but a quality score of 20 reflects a base call accuracy of 99%. These probability values are the results from the base calling algorithm and depend on how much signal was captured for the base incorporation.
Looking back at our read:
OUTPUT
@SRR2584863.1 HWI-ST957:244:H73TDADXX:1:1101:4712:2181/1
TTCACATCCTGACCATTCAGTTGAGCAAAATAGTTCTTCAGTGCCTGTTTAACCGAGTCACGCAGGGGTTTTTGGGTTACCTGATCCTGAGAGTTAACGGTAGAAACGGTCAGTACGTCAGAATTTACGCGTTGTTCGAACATAGTTCTG
+
CCCFFFFFGHHHHJIJJJJIJJJIIJJJJIIIJJGFIIIJEDDFEGGJIFHHJIJJDECCGGEGIIJFHFFFACD:BBBDDACCCCAA@@CA@C>C3>@5(8&>C:9?8+89<4(:83825C(:A#########################we can now see that there is a range of quality scores, but that the
end of the sequence is very poor (# = a quality score of
2).
Exercise
What is the last read in the SRR2584863_1.fastq file?
How confident are you in this read?
OUTPUT
@SRR2584863.1553259 HWI-ST957:245:H73R4ADXX:2:2216:21048:100894/1
CTGCAATACCACGCTGATCTTTCACATGATGTAAGAAAAGTGGGATCAGCAAACCGGGTGCTGCTGTGGCTAGTTGCAGCAAACCATGCAGTGAACCCGCCTGTGCTTCGCTATAGCCGTGACTGATGAGGATCGCCGGAAGCCAGCCAA
+
CCCFFFFFHHHHGJJJJJJJJJHGIJJJIJJJJIJJJJIIIIJJJJJJJJJJJJJIIJJJHHHHHFFFFFEEEEEDDDDDDDDDDDDDDDDDCDEDDBDBDDBDDDDDDDDDBDEEDDDD7@BDDDDDD>AA>?B?<@BDD@BDC?BDA?This read has more consistent quality at its end than the first read that we looked at, but still has a range of quality scores, most of them high. We will look at variations in position-based quality in just a moment.
At this point, lets validate that all the relevant tools are installed. If you are using the AWS AMI then these should be preinstalled.
Callout
On NeSI, we load the tools by running a script that has the needed software and versions specified.
Modules
On NeSI, software is managed through modules which
enable many versions of the same program to co-exist. This particular
modload.sh script contains the commands that loads the
specific software modules needed for our analysis. It uses the
module load command which is then followed by the module
you wish to use. In the case of fastqc we use
module load FastQC. To find the modules for software
managed on NeSI you can use
module spider <program name>,
e.g. module spider fastqc will return the names of the
modules related to fastqc.
For more information about using modules on NeSI see their documentation.
BASH
$ fastqc -h
FastQC - A high throughput sequence QC analysis tool
SYNOPSIS
fastqc seqfile1 seqfile2 .. seqfileN
fastqc [-o output dir] [--(no)extract] [-f fastq|bam|sam]
[-c contaminant file] seqfile1 .. seqfileN
DESCRIPTION
FastQC reads a set of sequence files and produces from each one a quality
control report consisting of a number of different modules, each one of
which will help to identify a different potential type of problem in your
data.
If no files to process are specified on the command line then the program
will start as an interactive graphical application. If files are provided
on the command line then the program will run with no user interaction
required. In this mode it is suitable for inclusion into a standardised
analysis pipeline.
The options for the program as as follows:
-h --help Print this help file and exit
-v --version Print the version of the program and exit
-o --outdir Create all output files in the specified output directory.
Please note that this directory must exist as the program
will not create it. If this option is not set then the
output file for each sequence file is created in the same
directory as the sequence file which was processed.
--casava Files come from raw casava output. Files in the same sample
group (differing only by the group number) will be analysed
as a set rather than individually. Sequences with the filter
flag set in the header will be excluded from the analysis.
Files must have the same names given to them by casava
(including being gzipped and ending with .gz) otherwise they
will not be grouped together correctly.
--nano Files come from naopore sequences and are in fast5 format. In
this mode you can pass in directories to process and the program
will take in all fast5 files within those directories and produce
a single output file from the sequences found in all files.
--nofilter If running with --casava then don't remove read flagged by
casava as poor quality when performing the QC analysis.
--extract If set then the zipped output file will be uncompressed in
the same directory after it has been created. By default
this option will be set if fastqc is run in non-interactive
mode.
-j --java Provides the full path to the java binary you want to use to
launch fastqc. If not supplied then java is assumed to be in
your path.
--noextract Do not uncompress the output file after creating it. You
should set this option if you do not wish to uncompress
the output when running in non-interactive mode.
--nogroup Disable grouping of bases for reads >50bp. All reports will
show data for every base in the read. WARNING: Using this
option will cause fastqc to crash and burn if you use it on
really long reads, and your plots may end up a ridiculous size.
You have been warned!
-f --format Bypasses the normal sequence file format detection and
forces the program to use the specified format. Valid
formats are bam,sam,bam_mapped,sam_mapped and fastq
-t --threads Specifies the number of files which can be processed
simultaneously. Each thread will be allocated 250MB of
memory so you shouldn't run more threads than your
available memory will cope with, and not more than
6 threads on a 32 bit machine
-c Specifies a non-default file which contains the list of
--contaminants contaminants to screen overrepresented sequences against.
The file must contain sets of named contaminants in the
form name[tab]sequence. Lines prefixed with a hash will
be ignored.
-a Specifies a non-default file which contains the list of
--adapters adapter sequences which will be explicity searched against
the library. The file must contain sets of named adapters
in the form name[tab]sequence. Lines prefixed with a hash
will be ignored.
-l Specifies a non-default file which contains a set of criteria
--limits which will be used to determine the warn/error limits for the
various modules. This file can also be used to selectively
remove some modules from the output all together. The format
needs to mirror the default limits.txt file found in the
Configuration folder.
-k --kmers Specifies the length of Kmer to look for in the Kmer content
module. Specified Kmer length must be between 2 and 10. Default
length is 7 if not specified.
-q --quiet Supress all progress messages on stdout and only report errors.
-d --dir Selects a directory to be used for temporary files written when
generating report images. Defaults to system temp directory if
not specified.
BUGS
Any bugs in fastqc should be reported either to simon.andrews@babraham.ac.uk
or in www.bioinformatics.babraham.ac.uk/bugzilla/if fastqc is not installed then you would expect to see an error like
$ fastqc -h
The program 'fastqc' is currently not installed. You can install it by typing:
sudo apt-get install fastqcIf this happens check with your instructor before trying to install it.
Assessing quality using FastQC
In real life, you will not be assessing the quality of your reads by visually inspecting your FASTQ files. Rather, you will be using a software program to assess read quality and filter out poor quality reads. We will first use a program called FastQC to visualize the quality of our reads. Later in our workflow, we will use another program to filter out poor quality reads.
FastQC has a number of features which can give you a quick impression of any problems your data may have, so you can take these issues into consideration before moving forward with your analyses. Rather than looking at quality scores for each individual read, FastQC looks at quality collectively across all reads within a sample. The image below shows one FastQC-generated plot that indicates a very high quality sample:
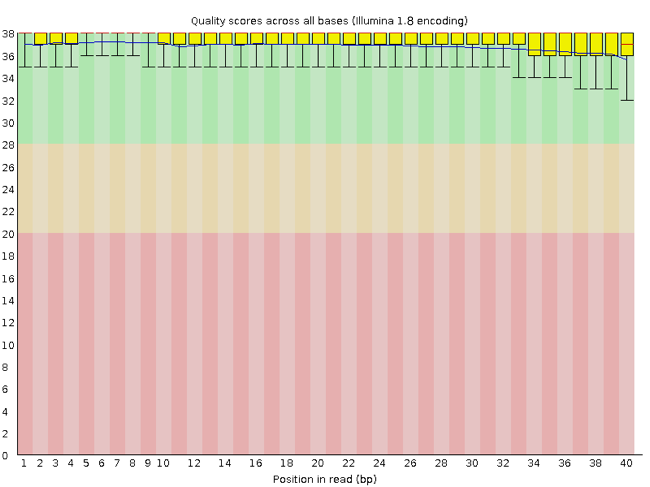
The x-axis displays the base position in the read, and the y-axis shows quality scores. In this example, the sample contains reads that are 40 bp long. This is much shorter than the reads we are working with in our workflow. For each position, there is a box-and-whisker plot showing the distribution of quality scores for all reads at that position. The horizontal red line indicates the median quality score and the yellow box shows the 1st to 3rd quartile range. This means that 50% of reads have a quality score that falls within the range of the yellow box at that position. The whiskers show the absolute range, which covers the lowest (0th quartile) to highest (4th quartile) values.
For each position in this sample, the quality values do not drop much lower than 32. This is a high quality score. The plot background is also color-coded to identify good (green), acceptable (yellow), and bad (red) quality scores.
Now let’s take a look at a quality plot on the other end of the spectrum.
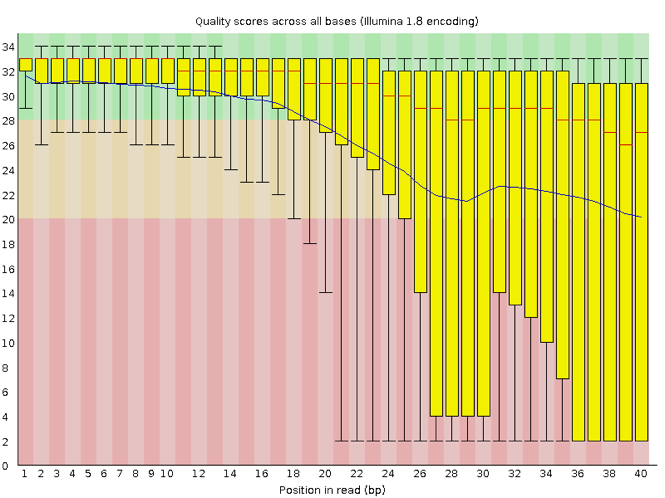
Here, we see positions within the read in which the boxes span a much wider range. Also, quality scores drop quite low into the “bad” range, particularly on the tail end of the reads. The FastQC tool produces several other diagnostic plots to assess sample quality, in addition to the one plotted above.
Running FastQC
We will now assess the quality of the reads that we downloaded.
First, make sure you are still in the untrimmed_fastq
directory
Exercise
How big are the files? (Hint: Look at the options for the
ls command to see how to show file sizes.)
OUTPUT
-rw-rw-r-- 1 dcuser dcuser 545M Jul 6 20:27 SRR2584863_1.fastq
-rw-rw-r-- 1 dcuser dcuser 183M Jul 6 20:29 SRR2584863_2.fastq.gz
-rw-rw-r-- 1 dcuser dcuser 309M Jul 6 20:34 SRR2584866_1.fastq.gz
-rw-rw-r-- 1 dcuser dcuser 296M Jul 6 20:37 SRR2584866_2.fastq.gz
-rw-rw-r-- 1 dcuser dcuser 124M Jul 6 20:22 SRR2589044_1.fastq.gz
-rw-rw-r-- 1 dcuser dcuser 128M Jul 6 20:24 SRR2589044_2.fastq.gzThere are six FASTQ files ranging from 124M (124MB) to 545M.
FastQC can accept multiple file names as input, and on both zipped and unzipped files, so we can use the *.fastq* wildcard to run FastQC on all of the FASTQ files in this directory.
You will see an automatically updating output message telling you the progress of the analysis. It will start like this:
OUTPUT
Started analysis of SRR2584863_1.fastq
Approx 5% complete for SRR2584863_1.fastq
Approx 10% complete for SRR2584863_1.fastq
Approx 15% complete for SRR2584863_1.fastq
Approx 20% complete for SRR2584863_1.fastq
Approx 25% complete for SRR2584863_1.fastq
Approx 30% complete for SRR2584863_1.fastq
Approx 35% complete for SRR2584863_1.fastq
Approx 40% complete for SRR2584863_1.fastq
Approx 45% complete for SRR2584863_1.fastqIn total, it should take about five minutes for FastQC to run on all six of our FASTQ files. When the analysis completes, your prompt will return. So your screen will look something like this:
OUTPUT
Approx 80% complete for SRR2589044_2.fastq.gz
Approx 85% complete for SRR2589044_2.fastq.gz
Approx 90% complete for SRR2589044_2.fastq.gz
Approx 95% complete for SRR2589044_2.fastq.gz
Analysis complete for SRR2589044_2.fastq.gz
$The FastQC program has created several new files within our
data/untrimmed_fastq/ directory.
OUTPUT
SRR2584863_1.fastq SRR2584866_1_fastqc.html SRR2589044_1_fastqc.html
SRR2584863_1_fastqc.html SRR2584866_1_fastqc.zip SRR2589044_1_fastqc.zip
SRR2584863_1_fastqc.zip SRR2584866_1.fastq.gz SRR2589044_1.fastq.gz
SRR2584863_2_fastqc.html SRR2584866_2_fastqc.html SRR2589044_2_fastqc.html
SRR2584863_2_fastqc.zip SRR2584866_2_fastqc.zip SRR2589044_2_fastqc.zip
SRR2584863_2.fastq.gz SRR2584866_2.fastq.gz SRR2589044_2.fastq.gzFor each input FASTQ file, FastQC has created a .zip
file and a
.html file. The .zip file extension
indicates that this is actually a compressed set of multiple output
files. We will be working with these output files soon. The
.html file is a stable webpage displaying the summary
report for each of our samples.
We want to keep our data files and our results files separate, so we
will move these output files into a new directory within our
results/ directory.
BASH
$ mkdir -p ~/obss_2024/genomic_dna/results/fastqc_untrimmed_reads
$ mv *.zip ~/obss_2024/genomic_dna/results/fastqc_untrimmed_reads/
$ mv *.html ~/obss_2024/genomic_dna/results/fastqc_untrimmed_reads/Now we can navigate into this results directory and do some closer inspection of our output files.
Viewing the FastQC results
We can use the explorer as part of Jupyter hub to navigate to the
directory containing the .html files and click on each one
to view the results of Fastqc.
Original intstuctions for AWS
If we were working on our local computers, we would be able to look at each of these HTML files by opening them in a web browser.
However, these files are currently sitting on our remote AWS instance, where our local computer can not see them. And, since we are only logging into the AWS instance via the command line - it does not have any web browser setup to display these files either.
So the easiest way to look at these webpage summary reports will be to transfer them to our local computers (i.e. your laptop).
To transfer a file from a remote server to our own machines, we will
use scp, which we learned yesterday in the Shell Genomics
lesson.
First we will make a new directory on our computer to store the HTML files we are transferring. Let’s put it on our desktop for now. Open a new tab in your terminal program (you can use the pull down menu at the top of your screen or the Cmd+t keyboard shortcut) and type:
Now we can transfer our HTML files to our local computer using
scp.
BASH
$ scp dcuser@ec2-34-238-162-94.compute-1.amazonaws.com:~/dc_workshop/results/fastqc_untrimmed_reads/*.html ~/Desktop/fastqc_htmlNote on using zsh
If you are using zsh instead of bash (macOS for example changed the
default recently to zsh), it is likely that a
no matches found error will be displayed. The reason for
this is that the wildcard (“*”) is not correctly interpreted. To fix
this problem the wildcard needs to be escaped with a “\”:
BASH
$ scp dcuser@ec2-34-238-162-94.compute-1.amazonaws.com:~/dc_workshop/results/fastqc_untrimmed_reads/\*.html ~/Desktop/fastqc_htmlAlternatively, you can put the whole path into quotation marks:
As a reminder, the first part of the command
dcuser@ec2-34-238-162-94.compute-1.amazonaws.com is the
address for your remote computer. Make sure you replace everything after
dcuser@ with your instance number (the one you used to log
in).
The second part starts with a : and then gives the
absolute path of the files you want to transfer from your remote
computer. Do not forget the :. We used a wildcard
(*.html) to indicate that we want all of the HTML
files.
The third part of the command gives the absolute path of the location
you want to put the files. This is on your local computer and is the
directory we just created ~/Desktop/fastqc_html.
You should see a status output like this:
OUTPUT
SRR2584863_1_fastqc.html 100% 249KB 152.3KB/s 00:01
SRR2584863_2_fastqc.html 100% 254KB 219.8KB/s 00:01
SRR2584866_1_fastqc.html 100% 254KB 271.8KB/s 00:00
SRR2584866_2_fastqc.html 100% 251KB 252.8KB/s 00:00
SRR2589044_1_fastqc.html 100% 249KB 370.1KB/s 00:00
SRR2589044_2_fastqc.html 100% 251KB 592.2KB/s 00:00Now we can go to our new directory and open the 6 HTML files.
Depending on your system, you should be able to select and open them all at once via a right click menu in your file browser.
Exercise
Discuss your results with a neighbor. Which sample(s) looks the best in terms of per base sequence quality? Which sample(s) look the worst?
All of the reads contain usable data, but the quality decreases toward the end of the reads.
Decoding the other FastQC outputs
We have now looked at quite a few “Per base sequence quality” FastQC graphs, but there are nine other graphs that we have not talked about! Below we have provided a brief overview of interpretations for each of these plots. For more information, please see the FastQC documentation here
- Per tile sequence quality: the machines that perform sequencing are divided into tiles. This plot displays patterns in base quality along these tiles. Consistently low scores are often found around the edges, but hot spots can also occur in the middle if an air bubble was introduced at some point during the run.
- Per sequence quality scores: a density plot of quality for all reads at all positions. This plot shows what quality scores are most common.
- Per base sequence content: plots the proportion of each base position over all of the reads. Typically, we expect to see each base roughly 25% of the time at each position, but this often fails at the beginning or end of the read due to quality or adapter content.
- Per sequence GC content: a density plot of average GC content in each of the reads.
- Per base N content: the percent of times that ‘N’ occurs at a position in all reads. If there is an increase at a particular position, this might indicate that something went wrong during sequencing.
- Sequence Length Distribution: the distribution of sequence lengths of all reads in the file. If the data is raw, there is often on sharp peak, however if the reads have been trimmed, there may be a distribution of shorter lengths.
- Sequence Duplication Levels: A distribution of duplicated sequences. In sequencing, we expect most reads to only occur once. If some sequences are occurring more than once, it might indicate enrichment bias (e.g. from PCR). If the samples are high coverage (or RNA-seq or amplicon), this might not be true.
- Overrepresented sequences: A list of sequences that occur more frequently than would be expected by chance.
- Adapter Content: a graph indicating where adapater sequences occur in the reads.
- K-mer Content: a graph showing any sequences which may show a positional bias within the reads.
Working with the FastQC text output
Now that we have looked at our HTML reports to get a feel for the
data, let’s look more closely at the other output files. Go back to the
tab in your terminal program that is connected to your AWS instance (the
tab label will start with dcuser@ip) and make sure you are
in our results subdirectory.
OUTPUT
SRR2584863_1_fastqc.html SRR2584866_1_fastqc.html SRR2589044_1_fastqc.html
SRR2584863_1_fastqc.zip SRR2584866_1_fastqc.zip SRR2589044_1_fastqc.zip
SRR2584863_2_fastqc.html SRR2584866_2_fastqc.html SRR2589044_2_fastqc.html
SRR2584863_2_fastqc.zip SRR2584866_2_fastqc.zip SRR2589044_2_fastqc.zipOur .zip files are compressed files. They each contain
multiple different types of output files for a single input FASTQ file.
To view the contents of a .zip file, we can use the program
unzip to decompress these files. Let’s try doing them all
at once using a wildcard.
OUTPUT
Archive: SRR2584863_1_fastqc.zip
caution: filename not matched: SRR2584863_2_fastqc.zip
caution: filename not matched: SRR2584866_1_fastqc.zip
caution: filename not matched: SRR2584866_2_fastqc.zip
caution: filename not matched: SRR2589044_1_fastqc.zip
caution: filename not matched: SRR2589044_2_fastqc.zipThis did not work. We unzipped the first file and then got a warning
message for each of the other .zip files. This is because
unzip expects to get only one zip file as input. We could
go through and unzip each file one at a time, but this is very time
consuming and error-prone. Someday you may have 500 files to unzip!
A more efficient way is to use a for loop like we
learned in the Shell Genomics lesson to iterate through all of our
.zip files. Let’s see what that looks like and then we will
discuss what we are doing with each line of our loop.
In this example, the input is six filenames (one filename for each of
our .zip files). Each time the loop iterates, it will
assign a file name to the variable filename and run the
unzip command. The first time through the loop,
$filename is SRR2584863_1_fastqc.zip. The
interpreter runs the command unzip on
SRR2584863_1_fastqc.zip. For the second iteration,
$filename becomes SRR2584863_2_fastqc.zip.
This time, the shell runs unzip on
SRR2584863_2_fastqc.zip. It then repeats this process for
the four other .zip files in our directory.
When we run our for loop, you will see output that
starts like this:
OUTPUT
Archive: SRR2589044_2_fastqc.zip
creating: SRR2589044_2_fastqc/
creating: SRR2589044_2_fastqc/Icons/
creating: SRR2589044_2_fastqc/Images/
inflating: SRR2589044_2_fastqc/Icons/fastqc_icon.png
inflating: SRR2589044_2_fastqc/Icons/warning.png
inflating: SRR2589044_2_fastqc/Icons/error.png
inflating: SRR2589044_2_fastqc/Icons/tick.png
inflating: SRR2589044_2_fastqc/summary.txt
inflating: SRR2589044_2_fastqc/Images/per_base_quality.png
inflating: SRR2589044_2_fastqc/Images/per_tile_quality.png
inflating: SRR2589044_2_fastqc/Images/per_sequence_quality.png
inflating: SRR2589044_2_fastqc/Images/per_base_sequence_content.png
inflating: SRR2589044_2_fastqc/Images/per_sequence_gc_content.png
inflating: SRR2589044_2_fastqc/Images/per_base_n_content.png
inflating: SRR2589044_2_fastqc/Images/sequence_length_distribution.png
inflating: SRR2589044_2_fastqc/Images/duplication_levels.png
inflating: SRR2589044_2_fastqc/Images/adapter_content.png
inflating: SRR2589044_2_fastqc/fastqc_report.html
inflating: SRR2589044_2_fastqc/fastqc_data.txt
inflating: SRR2589044_2_fastqc/fastqc.foThe unzip program is decompressing the .zip
files and creating a new directory (with subdirectories) for each of our
samples, to store all of the different output that is produced by
FastQC. There
are a lot of files here. The one we are going to focus on is the
summary.txt file.
If you list the files in our directory now you will see:
SRR2584863_1_fastqc SRR2584866_1_fastqc SRR2589044_1_fastqc
SRR2584863_1_fastqc.html SRR2584866_1_fastqc.html SRR2589044_1_fastqc.html
SRR2584863_1_fastqc.zip SRR2584866_1_fastqc.zip SRR2589044_1_fastqc.zip
SRR2584863_2_fastqc SRR2584866_2_fastqc SRR2589044_2_fastqc
SRR2584863_2_fastqc.html SRR2584866_2_fastqc.html SRR2589044_2_fastqc.html
SRR2584863_2_fastqc.zip SRR2584866_2_fastqc.zip SRR2589044_2_fastqc.zipThe .html files and the uncompressed .zip
files are still present, but now we also have a new directory for each
of our samples. We can see for sure that it is a directory if we use the
-F flag for ls.
OUTPUT
SRR2584863_1_fastqc/ SRR2584866_1_fastqc/ SRR2589044_1_fastqc/
SRR2584863_1_fastqc.html SRR2584866_1_fastqc.html SRR2589044_1_fastqc.html
SRR2584863_1_fastqc.zip SRR2584866_1_fastqc.zip SRR2589044_1_fastqc.zip
SRR2584863_2_fastqc/ SRR2584866_2_fastqc/ SRR2589044_2_fastqc/
SRR2584863_2_fastqc.html SRR2584866_2_fastqc.html SRR2589044_2_fastqc.html
SRR2584863_2_fastqc.zip SRR2584866_2_fastqc.zip SRR2589044_2_fastqc.zipLet’s see what files are present within one of these output directories.
OUTPUT
fastqc_data.txt fastqc.fo fastqc_report.html Icons/ Images/ summary.txtUse less to preview the summary.txt file
for this sample.
OUTPUT
PASS Basic Statistics SRR2584863_1.fastq
PASS Per base sequence quality SRR2584863_1.fastq
PASS Per tile sequence quality SRR2584863_1.fastq
PASS Per sequence quality scores SRR2584863_1.fastq
WARN Per base sequence content SRR2584863_1.fastq
WARN Per sequence GC content SRR2584863_1.fastq
PASS Per base N content SRR2584863_1.fastq
PASS Sequence Length Distribution SRR2584863_1.fastq
PASS Sequence Duplication Levels SRR2584863_1.fastq
PASS Overrepresented sequences SRR2584863_1.fastq
WARN Adapter Content SRR2584863_1.fastqThe summary file gives us a list of tests that FastQC ran, and tells
us whether this sample passed, failed, or is borderline
(WARN). Remember, to quit from less you must
type q.
Documenting our work
We can make a record of the results we obtained for all our samples
by concatenating all of our summary.txt files into a
single file using the cat command. We will call this
fastqc_summaries.txt and move it to
~/obss_2024/genomic_dna/docs.
Exercise
Which samples failed at least one of FastQC’s quality tests? What test(s) did those samples fail?
We can get the list of all failed tests using grep.
OUTPUT
FAIL Per base sequence quality SRR2584863_2.fastq.gz
FAIL Per tile sequence quality SRR2584863_2.fastq.gz
FAIL Per base sequence content SRR2584863_2.fastq.gz
FAIL Per base sequence quality SRR2584866_1.fastq.gz
FAIL Per base sequence content SRR2584866_1.fastq.gz
FAIL Adapter Content SRR2584866_1.fastq.gz
FAIL Adapter Content SRR2584866_2.fastq.gz
FAIL Adapter Content SRR2589044_1.fastq.gz
FAIL Per base sequence quality SRR2589044_2.fastq.gz
FAIL Per tile sequence quality SRR2589044_2.fastq.gz
FAIL Per base sequence content SRR2589044_2.fastq.gz
FAIL Adapter Content SRR2589044_2.fastq.gzOther notes – optional
Quality encodings vary
Although we have used a particular quality encoding system to
demonstrate interpretation of read quality, different sequencing
machines use different encoding systems. This means that, depending on
which sequencer you use to generate your data, a # may not
be an indicator of a poor quality base call.
This mainly relates to older Solexa/Illumina data, but it is essential that you know which sequencing platform was used to generate your data, so that you can tell your quality control program which encoding to use. If you choose the wrong encoding, you run the risk of throwing away good reads or (even worse) not throwing away bad reads!
Same symbols, different meanings
Here we see > being used as a shell prompt, whereas
> is also used to redirect output. Similarly,
$ is used as a shell prompt, but, as we saw earlier, it is
also used to ask the shell to get the value of a variable.
If the shell prints > or $ then
it expects you to type something, and the symbol is a prompt.
If you type > or $ yourself, it
is an instruction from you that the shell should redirect output or get
the value of a variable.
Key Points
- Quality encodings vary across sequencing platforms.
-
forloops let you perform the same set of operations on multiple files with a single command.
